HeartVolMesh
Cardiac Volumetric Mesh Reconstruction via Covariance-Guided Graph Deformation
Watch the HeartVolMesh introduction
The player opens with the English version selected by default. Use the buttons below to switch between English and Chinese while keeping the native playback controls for seeking, pausing, volume, and fullscreen. This version does not start muted.
From 3D CTA volumes to simulation-ready tetrahedral heart meshes: HeartVolMesh combines covariance-guided graph deformation with template-driven volumetric warping to preserve topology, correspondence, and downstream simulation utility.
Why this matters
In-silico cardiovascular studies need more than a surface boundary. They need a patient-specific volumetric mesh that can be used by FEM/CFD solvers, while remaining faithful around thin walls and multi-structure junctions.
Core idea
HeartVolMesh lifts each template vertex from a deterministic point to an anisotropic Gaussian kernel, predicts both displacement and covariance, and then warps a tetrahedral template to the reconstructed target surface.
What this gives
A topology-consistent surface plus a correspondence-preserving tetrahedral mesh, with resolution controlled by the chosen template instead of per-case tetrahedralization.
Abstract
Accurate patient-specific tetrahedral cardiac meshes are essential for in-silico trials, yet common segmentation-then-modelling pipelines can blur thin-wall anatomy and offer limited cross-case correspondence. HeartVolMesh addresses this with two complementary ideas. First, it reconstructs a topology-consistent surface by predicting per-vertex displacements together with Cholesky-parameterized anisotropic covariances, so uncertainty and local geometric support are modelled directly during learning. Second, it warps a fixed tetrahedral template to the reconstructed surface through staged alignment, non-rigid registration, and deformation-field propagation, preserving connectivity and correspondence by construction. The result is a simulation-oriented image-to-mesh pipeline that improves surface accuracy while unlocking high-quality volumetric reconstruction.
Contributions
Covariance-guided graph deformation
Each template vertex is represented as an anisotropic Gaussian kernel, allowing the network to learn direction-aware geometric tolerance rather than relying only on uniform Euclidean supervision.
Template-driven volumetric meshing
A high-quality tetrahedral template is aligned to the predicted surface and deformed through a dense deformation field, preserving mesh connectivity and enabling cross-case correspondence.
Whole-heart validation
The framework is evaluated on multi-structure cardiac reconstruction and shows consistent gains in both surface-mesh accuracy and volumetric boundary fidelity, together with strong element quality.
Representation and learning objective
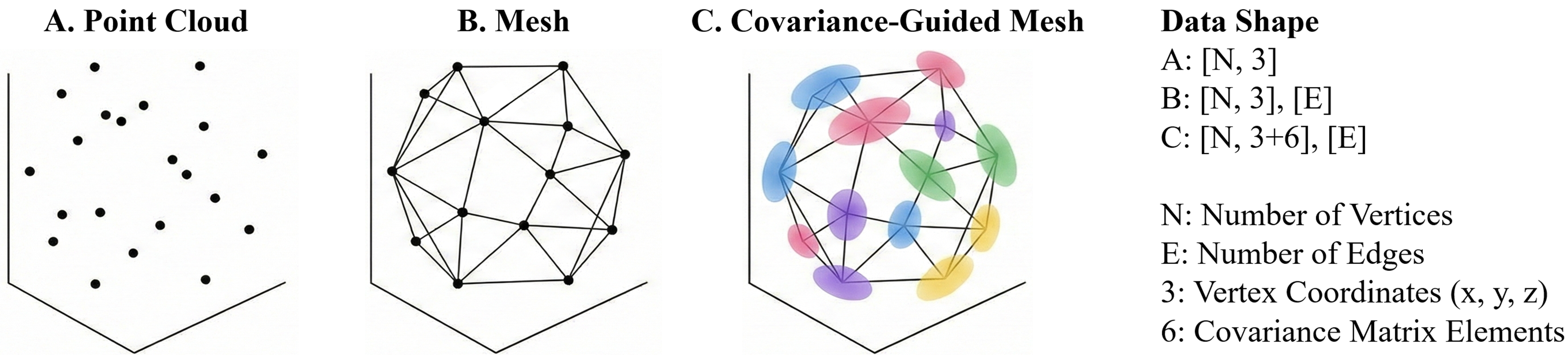
From points to covariance-guided meshes
- Classical point sets store isolated coordinates.
- Classical meshes add graph connectivity but still treat each vertex as a point.
-
HeartVolMesh extends every vertex to
[x, y, z] + 6covariance parameters, allowing local anisotropy to be learned directly from the volume.
Covariance-guided matching loss
Surface supervision uses a covariance-aware negative log-likelihood objective with lightweight edge, Laplacian, and normal regularization:
$$ \ell(x, y; \Sigma)=\frac{1}{2}(x-y)^\top \Sigma^{-1}(x-y)+\beta\log\det(\Sigma). $$This lets the model express spatially varying and direction-dependent uncertainty, which is especially useful near thin walls and multi-structure junctions.
Pipeline at a glance
Image-conditioned vertex features
A 3D CNN encoder extracts multi-scale volumetric features, which are sampled at current vertex positions and propagated over the mesh graph with a GNN.
Coarse-to-fine surface prediction
Meshes are progressively refined across stages. The final head predicts both vertex displacements and Cholesky parameters for covariance construction.
Surface-to-volume propagation
Surface motion is converted into a dense deformation field and applied to every vertex of a tetrahedral template, yielding a volumetric mesh whose boundary matches the reconstructed patient-specific surface.
Experimental setting
Dataset and training
- Multi-centre in-house dataset with 900 patients and 4,000 temporal instances.
- Patient-level split: 800 patients / 3,600 instances for training and 100 patients / 400 instances for validation.
- Single NVIDIA A100 80GB, Adam optimizer, 100 epochs, batch size 1.
- Input crops of
128^3; deformation field predicted at300^3.
Evaluation protocol
- Surface reconstruction on LA, LV, RA, RV, and LVMyo.
- Volumetric reconstruction on LAMyo, LVMyo, RAMyo, and RVMyo.
- Boundary metrics: symmetric Chamfer Distance, HD95, and Normal Consistency.
- Volumetric quality: inverted elements, minimum scaled Jacobian, and minimum dihedral angle.
Qualitative volumetric reconstruction
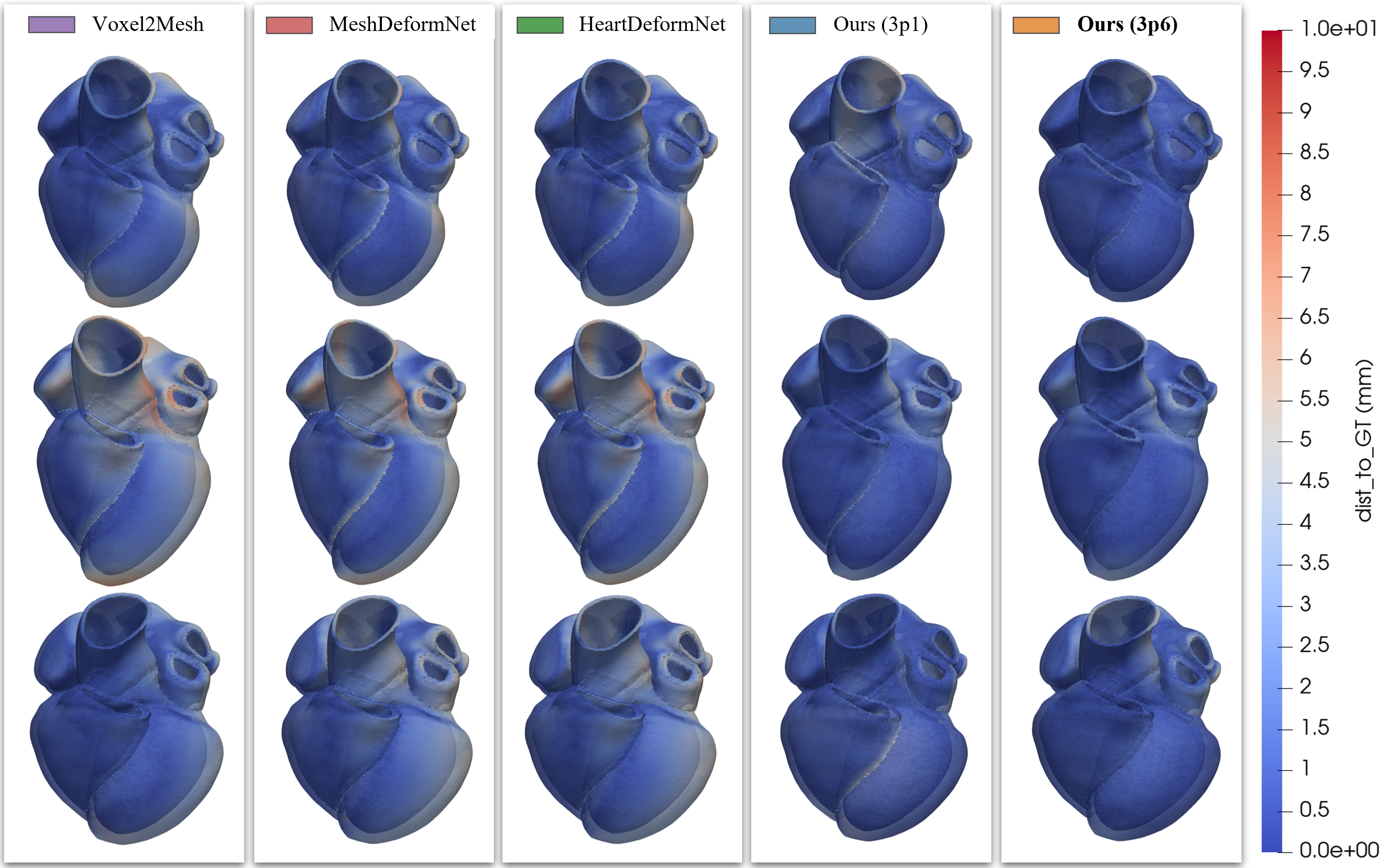

Quantitative summary
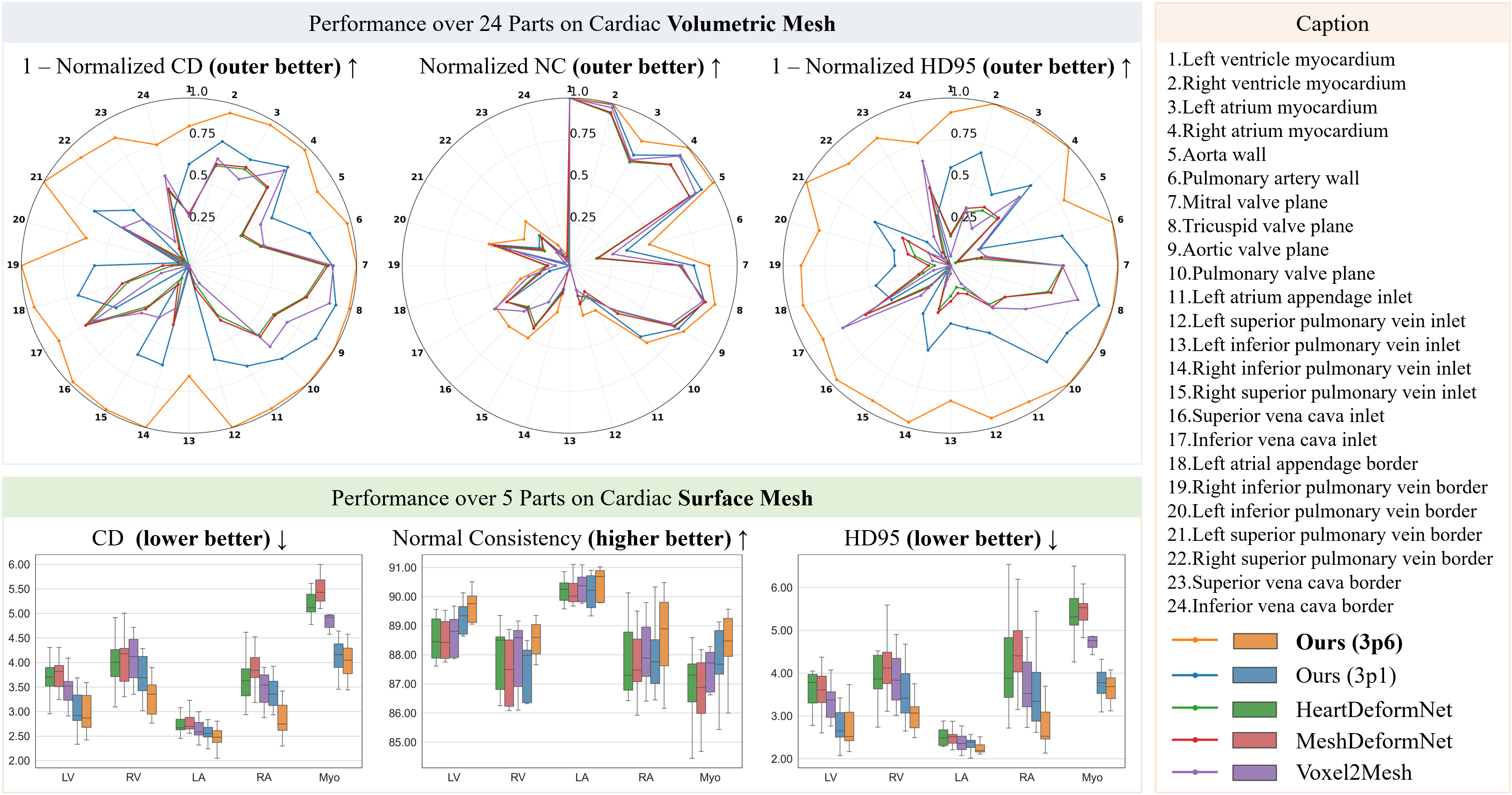
Best surface-mesh CD / HD95 / NC (Ours 3p6)
| Structure | CD ↓ | HD95 ↓ | NC ↑ |
|---|---|---|---|
| LA | 2.5 | 2.2 | 89.5% |
| LV | 3.0 | 2.8 | 88.7% |
| Myo | 4.0 | 3.6 | 87.6% |
| RA | 2.9 | 2.8 | 88.3% |
| RV | 3.3 | 3.1 | 87.3% |
Best volumetric-boundary CD / HD95 / NC (Ours 3p6)
| Structure | CD ↓ | HD95 ↓ | NC ↑ |
|---|---|---|---|
| LVMyo | 3.6 | 3.2 | 95.3% |
| RVMyo | 3.1 | 2.8 | 94.2% |
| LAMyo | 3.0 | 3.0 | 87.7% |
| RAMyo | 3.0 | 2.9 | 92.2% |
Key takeaways
Better boundary localization
Covariance-guided supervision improves robustness where uniform Euclidean penalties are too rigid, especially at ambiguous and thin anatomical boundaries.
Volumetric meshes with correspondence
By deforming a fixed tetrahedral template instead of tetrahedralizing each case independently, the framework preserves connectivity and enables cross-case vertex correspondence by design.
Simulation-oriented flexibility
The method trades fully end-to-end meshing for a template-conditioned formulation that can support different mesh densities and element specifications when an appropriate template is provided.
Citation
1
2
3
4
5
6
7
8
@inproceedings{heartvolmesh2026,
title = {HeartVolMesh: Cardiac Volumetric Mesh Reconstruction via Covariance-Guided Graph Deformation},
author = {Anonymous},
booktitle = {ArXiv preprint},
year = {2026},
note = {Review version; replace with final metadata after de-anonymization}
}
NVIDIA Academic Grant Support
This project was made possible by the NVIDIA Academic Grant Program
HeartVolMesh was made possible by the same NVIDIA Academic Grant support that underpinned our broader research effort, even though the final direction of this project evolved beyond the scope of the original proposal.
This support helped accelerate the compute-intensive experimentation behind the project—from 3D image-conditioned surface reconstruction to template-driven volumetric warping and validation. Learn more about the NVIDIA Academic Grant Program.
NVIDIA-supported academic compute helped accelerate development, experimentation, and validation for this work.